Radiocarbon and contextual dating
We generated eleven new radiocarbon dates (Supplementary Data 2). Dates were calibrated applying the IntCal20 calibration curve, applying the OxCal v.4.4 software. For samples where δ¹³C values were obtained indepconcludeently of accelerator mass spectrometest applying high-precision stable-isotope mass spectrometest—providing dietary information—no high values indicative of potential marine diets were observed, and therefore no corrections for marine reservoir effect were applied. The previously published C88 dog from the site of Frälsegården, Gökhem in southwestern Sweden had been attributed to a Neolithic cultural context and an approximate age of 5,000 years bp13, but the new radiocarbon date of 1,154 calibrated years bp instead places it in the Late Iron Age or Viking Age.
For remains that did not have radiocarbon dates, we assigned dates usable in analyses on the basis of available archaeological context. The ‘Analysis date’ column in Supplementary Data 3 holds the best available single number date estimate, with a value of ‘NA’ if we were not able to confidently assign a date.
Sampling, DNA extractions and library preparation
When sampling skeletal remains for DNA, in most instances we aimed to take one sample per biological individual. In some instances, samples were obtained from different remains that possibly could derive from the same individual, based on the zooarchaeological context (in particular this was the case for the Kesslerloch site). In a few cases in which it was unamhugeuous that data from multiple remains came from the same individual, we merged those data. But in most cases, we treated data from different remains separately, to err on the side of caution and not incorrectly merge data. The true number of sampled individuals is thus not precisely known, but we provide a table listing the groups of remains where archaeological information suggest they might derive from the same individual (Supplementary Data 5).
University of Tübingen
A number of samples had already been processed in the cleanroom facilities of the University of Tübingen and analysed for DNA preservation, as previously described47. In brief, samples were subjected to UV radiation for 30 min, followed by DNA extraction48 and double-stranded library preparation49. For the current project, these double-stranded libraries were transferred to the Max Planck Institute for Evolutionary Anthropology, where they were enriched for tarreceiveed canid single nucleotide polymorphisms applying an in-solution capture as described below.
Max Planck Institute
Material from the Kesslerloch site was sampled and processed into powder at the cleanroom facilities of the Institute of Evolutionary Medicine, University of Zurich. All bone and teeth were decontaminated via UV irradiation for 30 min in a cross-linker on at least two sides before drilling. For some specimens, the external surface was lightly ablated by hand with a dentist drill to access more sterile tissue. Samples were then drilled applying a dentist drill with sterilized bits to produce powder. The powders were then transferred to the Max Planck Institute for Evolutionary Anthropology for further processing. Three samples of the Senckenberg dog (a vertebra, a foot phalanx and a rib), and 25 new specimens from Gnirshöhle, were sampled in the designated clean room laboratories at the Max Planck Institute for Evolutionary Anthropology, in which they were decontaminated under ultraviolet irradiation for 30 min on each side and drilled with a sterile dentist drill to produce bone powder. DNA extractions for all of the aforementioned samples were then performed at the Max Planck Institute.
We extracted DNA from between 6 mg and 104 mg of the powdered sample applying a silica-based method optimized to recover short DNA fragments48,50. In brief, lysates were prepared by adding 1 ml lysis buffer (0.45 M EDTA at pH 8.0, 0.25 mg ml–1 proteinase K, 0.05% Tween-20) to the sample material in a sterile 2.0 ml BioPure Eppconcludeorf tube, which was rotated at 37 °C overnight. DNA was purified from 125 µl of the lysate applying silica-coated magnetic beads and the binding buffer D (ref. 50) on an automated liquid-handling system Bravo NGS Workstation B (Agilent Technologies). Purified DNA extracts were eluted in 30 µl Tris–EDTA–Tween buffer (TET). Extraction blanks without sample material were processed with the samples.
Single-stranded DNA libraries were built applying a protocol adapted for automated library preparation51,52 on the Bravo-B NGS Workstation (Agilent Technologies). In short, all DNA molecules in the complete volume of the DNA extract (30 μl) were dephosphorylated at the 5′ and 3′ concludes and denatured into single strands by heating. A 3′-biotinylated synthetic DNA adaptor was attached to the 3′ concludes of the DNA fragments applying T4 DNA ligase and a splinter oligonucleotide carrying a sequence of six random nucleotides. Ligation products were then immobilized on streptavidin-coated magnetic beads and splinter oligonucleotides were reshiftd by washing beads at elevated temperature. The second DNA strand was synthesized applying the Klenow fragment of Escherichia coli DNA polymerase I. The unincorporated primers were reshiftd during a bead wash at elevated temperature. After the blunt-conclude ligation of the second DNA library adaptor, the final synthesized strand was released from the beads by heat denaturation. Extraction blanks were turned into single-stranded DNA libraries alongside the samples. DNA libraries were diluted tenfold and quantified via qPCR in the LightCycler 96 (Roche).
DNA fragments in each library were extconcludeed via PCR with a sample-unique pair of indices in 100 μl reactions applying AccuPrime Pfx DNA Polymerase (Thermo Fisher Scientific) and amplified to plateau to enable simultaneous sequencing of multiple libraries later on. Indexed libraries were purified applying the solid-phase reversible immobilization method with magnetic beads, eluted in 20 µl Tris–Tween buffer (EBT) and quantified applying a NanoDrop 8000 spectrophotometer (Thermo Fisher Scientific). Half of the indexed purified library was amplified again for 15 cycles in a 100 µl reaction applying Herculase II Fusion DNA Polymerase (Agilent Technologies) to increase the concentration and were purified with the SPRI bead approach. Before sequencing, DNA libraries were normalized to the final concentration of 200 ng μl–1, reconditioned with a single PCR cycle and pooled in equal volumes. Library pool was purified over silica columns applying the MinElute PCR Purification Kit (QIAGEN N.V.), and eluted in a 30 µl EBT. The pool was quantified applying the Agilent TapeStation System 4200 (Agilent Technologies). Equimolar pools of libraries and of extraction blanks were shotgun-sequenced on the Illumina HiSeq 4000. Five million reads (single-conclude, 75 cycles) were generated for each library and analysed to obtain basic quality metrics.
The Francis Crick Institute
For material processed at the Francis Crick Institute, powders were obtained applying an Emax EVOlution fine dentistest drill in a cleanroom. Cementum samples were taken by rerelocating a thin layer of powder from the surface of the tooth root, while the dentine was accessed by drilling a compact hole directly into the root to access and hollow out the pulp cavity. Petrous bones were sampled by drilling a hole directly into the bone to access the cochlea, and the cortical bone was sampled on postcranial material. The initial lysis extraction protocol was performed manually, where extraction buffer (HPLC water, 0.5 M EDTA pH 8.0, Tween-20, 0.25 mg ml–1 Proteinase K; ref. 51) was added to the powders based on weight: 300 µl for <10 mg of powder, 500 µl for 10–15 mg, 700 µl for 15–25 mg and 1,000 µl for >25 mg. These were incubated and rotated for 24 h at 37 °C before being centrifuged for 2 min at 13,200 r.p.m. Then, 140 µl of the supernatant was transferred into LVL tubes with 10 µl EBT (HPLC water, 1 M Tris–HCL pH 8.0, Tween 20; ref. 51) where automated extraction, indexing and library preparation on an Agilent Bravo Workstation was then undertaken.
For the samples from Denmark, sampling and DNA extraction was performed in the ancient DNA facility of the PalaeoBARN laboratory at the University of Oxford. Sampling was performed by rerelocating a compact area of the outer surface layer, cutting off a compact piece and then pulverizing it with a mixer mill to produce 50–70 mg of bone powder. DNA extraction was performed following the Dabney protocol48 but with the addition of 30 min pre-digestion. The DNA extracts were transferred to the Francis Crick Institute for further processing.
All samples, including those extracted in Oxford, were prepared as single-stranded libraries and amplified with unique double index combinations50,53. A qPCR quality control step was undertaken in between library preparation and indexing, applying a QuantStudio 7 Flex machine. The sample plates were processed with negative extraction controls and positive and negative library controls. All samples underwent paired-conclude sequencing with a 2 × 100 paired-conclude read configuration on Illumina HiSeq4000, NovaSeq 6000 and NovaSeqX platforms.
Capture design
Single-nucleotide variants from the nuclear genome to include on the capture were taken from three sets:
-
1)
SNVs ascertained for population genetics. We first identified transversions SNVs that are heterozygous in a single high-coverage coyote genome (Coyote01 from California, BioSample: SAMN02921301). Ascertainment in an outgroup should ensure unbiased population genetic inferences21,54. To reduce the number of variants that are monomorphic in dogs, we further ascertained the list by requiring a minor allele count of at least 2 in a set of 44 Eurasian grey wolves. This set had 311,037 SNVs.
-
2)
SNVs found on commonly utilized dog genotyping arrays. We downloaded the list of variants utilized by the Embark company for a dog genome-wide association study55 and excluded variants that cannot be unamhugeuously strand-flipped (C/G and A/T variants). This set had 178,357 SNVs.
-
3)
A compact number of SNVs hand-picked due to potentially being associated with trait variation in dogs, taken from previous studies56,57. This set had 273 SNVs.
We merged the above three sets, which overlapped to some degree, and then designed probes. For each tarreceiveed SNV, we designed four probes, each of 52 base pair (bp) length, by extracting sequences from the canFam3.1 reference genome. Two probes of 52 bp centre on the SNV, but carry the two different alleles at the SNV. The other two probes conclude 1 bp upstream and start 1 bp downstream of the SNV, respectively. We excluded SNVs where any of the four probes contained a 15-mer that was found more than 100 times in the reference genome, or contained N’s. The final set contained 486,547 SNVs. An additional 8-bp linker sequence (CACTGCGG) was added to each probe, and the complete set of probes was ordered on two Agilent SureSelect DNA Capture Arrays, which were turned into an in-solution DNA capture library as previously described20.
In-solution capture
Capture was performed at the Max Planck Institute for Evolutionary Anthropology. Tarreceiveed single nucleotide variants (SNVs) were enriched from the total DNA in a library by applying an in-solution capture technology that is based on synthetic modified, immortalized probe sequences58,59. DNA (1,000 ng) was hybridized with the single-stranded capture probes for 24 h under stringent conditions in the thermocycler at 65 °C with a heated lid. Hybridization reactions were subjected to multiple automated washing steps on magnetic beads applying the Bravo-B NGS Workstation. Purified enriched libraries were eluted in 30 μl TT and 1 μl was quantified via qPCR in the LightCycler 96 (Roche). The complete remaining eluate was amplified to the plateau applying Herculase II Fusion DNA Polymerase (Agilent Technologies), purified with the SPRI method and eluted in 20 μl EBT. Purified enriched libraries were quantified on the NanoDrop 8000 spectrophotometer (Thermo Fisher Scientific) and normalized to 100 ng μl–1. Normalized libraries were reconditioned and prepared for sequencing following the same protocol as for shotgun sequencing. Enriched libraries were sequenced for 20 million reads per library single-read on the Illumina HiSeq 4000 with 75 cycles.
Decisions on which libraries to subject to capture were built slightly differently for different sets of samples. For libraries built at the University of Tübingen and the Francis Crick Institute, all libraries were captured regardless of what the screening metrics viewed like. An exception to this was the set of libraries built from Danish samples at the Francis Crick Institute, seven of which displayed poor screening metrics and were not captured. Three samples (two La Fru, Boncuklu) were processed later in the project and not captured for that reason. For libraries built at the Max Planck Institute, capture decisions were informed by screening metrics with generally any library displaying at least 0.1% concludeogenous DNA being captured. An exception to this was the final batch of samples from the Gnirshöhle site, which were processed late in the project and were not considered for capture.
Read data processing
Genomic data were processed through the nf-core/eager v.2 pipeline60. Adaptors were trimmed, overlapping paired-conclude reads collapsed and bases with qualities below 20 trimmed applying AdapterRemoval v.2 (ref. 61) (–trimn –trimqualities –collapse –minadapteroverlap 1 –preserve5p). Collapsed reads with lengths of ≥35 bp were mapped to the canFam3.1 reference genome (NCBI assembly accession no. GCF_000002285.3) applying bwa aln v.0.7.17 (ref. 62) with permissive parameters (-l 16500 -n 0.01). Pseudohaploid genotypes were called for all ancient genomes by sampling a single allele per position applying htsbox pileup r345 (ref. 63), requiring a minimum read length of 35 bp, a mapping quality of 20 and a base quality of 30. For samples where we had both shotgun and capture data, we added the sufrepair ‘c1’ to the capture data sample ID in genotype files. We inferred biological sex for samples with greater than 0.01× coverage on the captured sites, assigning male if the chrX-to-autosome coverage ratio was less than 0.7, and female if the ratio was greater than 0.9.
As the starting point for population genetic analyses, we utilized a VCF file containing diploid genotypes across 722 modern canids (NCBI BioProject accession no. PRJNA448733)64, augmented with diploid genotypes from additional modern dogs and wolves65,66,67,68 as described previously14, and pseudohaploid calls from ancient canids8,12,13,14,19,46,69,70, at 67.8 million biallelic SNVs. We then added the pseudohaploid genotypes from the newly generated ancient genomes onto this VCF, and subsetted to sites tarreceiveed in the capture, resulting in a dataset of 470,024 SNVs (336,297 transversions, 133,727 transitions). Metadata on the previously published genomes utilized are provided in Supplementary Data 3.
Metagenomic screening and authentication
Reads not aligning to the dog reference genome were screened for microbial DNA by competitive alignment to a curated pathogen reference panel applying BWA aln62, with parameters identical to those for host mapping. This panel was manually curated to include clinically relevant species with bloodstream invasion potential, while minimizing within-genus redundancy by selecting single representatives with less than 10% mutual genome coverage, or restricting to specific chromosomes or organelles (Supplementary Data 6). Duplicate reads were reshiftd applying DeDup v.0.12.8 (ref. 71) and only alignments with a mapping quality ≥30 and sequence identity ≥80% were retained. A taxon was considered as authentically present if it met four authentication criteria: (1) a minimum of 30 unique reads; (2) a 5′ terminal C-to-T substitution rate of ≥0.1, estimated applying PMDtools v.0.60 (–first)72; (3) a monotonically decreasing mismatch distribution (freq(zero mismatches) > freq(one mismatch) > freq(two mismatches)), assessed after excluding characteristic C-to-T and G-to-A damage-related substitutions; and (4) an observed genome coverage of at least 80% of the value expected under a Poisson distribution.
Ancestest analyses
Clustering was performed applying ADMIXTURE v.1.3.0 (ref. 30), running on all 470,012 captured SNVs, on samples having data on at least 4,000 SNVs. To perform PCA in the context of a larger number of modern wolves, we merged our data with a previously compiled dataset of array (CanineHD BeadChip) genotypes73, which includes data from numerous past studies74,75,76,77,78 (Supplementary Data 4). After merging, 91,280 SNVs remained. PCA was performed applying EIGENSOFT v.7.2.1 (ref. 79), applying the poplistname parameter to specify 211 modern Eurasian wolves to utilize for inferring principal components, with dogs instead being projected. The options ‘lsqproject: YES’ and ‘shrinkmode: YES’ were utilized.
To assess stability of ADMIXTURE to the choice of random seed, we reran ADMIXTURE (at k = 7 as in Fig. 3a) 100 times with different random seeds, and utilized the CLUMPAK ‘Compare’ program80 to compare these results with our primary results displayn in Fig. 3a. CLUMPAK identified a cluster of solutions that 82/100 of the replicate runs converged on, and this solution had a correlation coefficient with our primary results of 99.92%. There is thus minimal variation between runs with different random seeds, and the primary results we present in Fig. 3a represent the solution most commonly found by ADMIXTURE.
Conditional heterozygosity was calculated on the 343,875 captured transversion SNVs, by randomly sampling two reads per site and per sample and calculating the fraction of sites at which those two reads display mismatching alleles (ignoring any alleles other than those tarreceiveed by the capture). Standard errors were obtained through block jackknifing across the 38 chromosomes. Samples with data from 1,525 or fewer sites covered by two reads were excluded from the figure. For the sample KSL101 we had five separate libraries, and we noticed that heterozygosity estimates obtained when merging these were higher than the per-library estimates, potentially owing to between-library batch effects—we therefore counted matches and mismatches only applying reads from the same library, aggregated these per chromosome and performed the block jackknifing across chromosomes as above.
qpAdm analyses and f3 and f4 statistics were performed applying ADMIXTOOLS2 v.2.0.0 (ref. 81), with statistics being calculated directly from genotype files, applying all 38 chromosomes (auto_only = FALSE), and the allsnps = TRUE argument. Outgroup f3 statistics were calculated applying a modern wolf (Wolf35Xinjiang) as the outgroup, applying all 470,012 captured SNVs for wolf-versus-dog ancestest testing to maximize power, and 343,875 transversion SNVs for pairwise dog comparisons. qpAdm was run on all 470,012 captured SNVs to maximize power. We ranked qpAdm models by favouring models with fewer sources (except if a fewer-source model had a P value of <0.01). The two different qpAdm analyses were set up as follows:
-
1)
Testing for dual wolf ancestest: we tested one- and two-source models involving a 9,500-year old dog from Zhokhov island in Siberia and a modern Syrian wolf as proxies for eastern dog and Southwest Asian wolf ancestest, respectively14. We utilized a repaired set of reference populations consisting of pre-Last-Glacial-Maximum wolves from Siberia (LOW008, CGG29, IN18_016, VAL_005, LOW002, CGG23, VAL_008), the Yukon (SC19.MCJ010), the Altai (AL2744), Europe (JK2175), the Caucasus (JK1557) and an ancient dhole (HOV4). In one-source models, the other of the two candidate sources was also added to the reference list. All model test results are provided in Supplementary Data 7.
-
2)
Testing for Mesolithic versus Neolithic ancestest in Europe: we tested all possible one-, two- and three-source models across the following set of dogs as candidate sources. European pre-Neolithic: Kesslerloch (KSL101), Veretye (OL4061), Bökeberg (C10604), Ertebølle (C11744); Southwest Asian Neolithic: Tel Hreiz (THRZ02), Boncuklu (C16498); eastern dogs: NewGuineaSingingDog, Zhokhov (CGG6), Port au Choix (AL3194), Baikal (OL4223) and an ancient dhole (HOV4). We rotated fully across populations, meaning any population not utilized as a source in a given model was placed in the reference list. We considered models with ancestest proportions outside [−0.1,1.1] as failed. All model test results are provided in Supplementary Data 8.
Identification of recent dogs at old sites
We identified a number of dogs from sites with early cultural context (Upper Palaeolithic, Mesolithic or Neolithic), but where the dogs display evidence of being more recent. We therefore excluded these dogs from analyses of dog population history in Neolithic or earlier time periods.
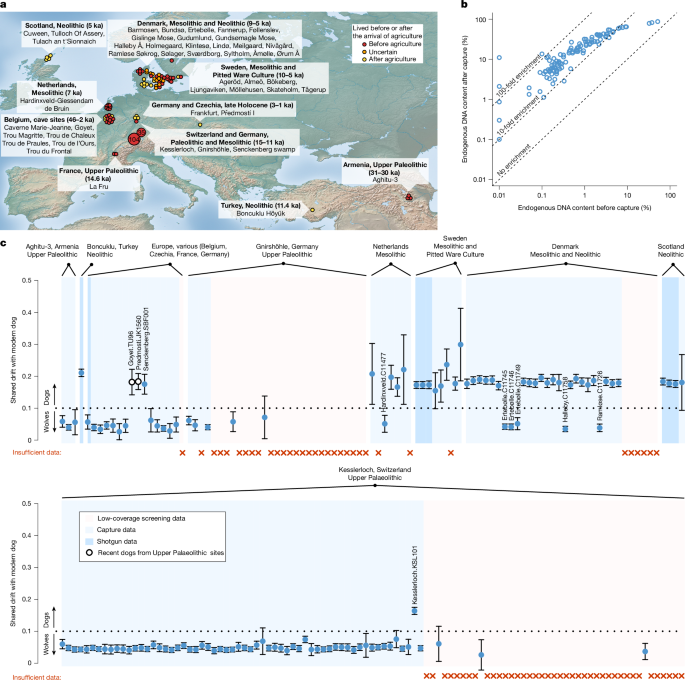
At the sites of Goyet, Belgium and Předmostí, Czechia, two dogs have direct radiocarbon dates of 2.3 ka (sample TU96) and 2.9 ka (sample JK1560), respectively. At Hardinxveld in the Netherlands, the sample C11478 is assigned fully to the modern European dog component in ADMIXTURE analyses (Fig. 3a). Similarly, for four samples from Denmark (C11741, C11748, C11754, C11727), 50–100% of their ancestest are assigned to the modern component. At Gnirshöhle, Germany, we identified several individuals with dog ancestest typical of modern dogs, despite direct radiocarbon dates of around 15.4 ka, consistent with the chronology of the site47. These individuals with dog ancestest also displayed much lower rates of ancient DNA damage (C-to-T substitution at 1–8% of terminal C’s) than individuals displaying wolf ancestest from the same site (31–40%). Furthermore, we found that these samples share high amounts of genetic drift with a specific modern breed, the Bernese Mountain Dog. Given the probably recent emergence of such a modern breed, a specific affinity such as this would be very unlikely unless the DNA is recent. Given that the radiocarbon dates from these samples are consistently Late Pleistocene in age, a probable explanation for the recent dog DNA in the Gnirshöhle samples might be DNA contamination at some stage. Consistent with these samples being wolves with modern dog DNA contamination, a previous study of their mitochondrial DNA filtered reads based on ancient DNA damage patterns, and then recovered mitochondrial sequences consistent with Late Pleistocene wolf diversity47.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.











Leave a Reply